Home » DataWarrior » Functionality » Macro for evolutionary library (EL) in Datwarrior (DW)
| Macro for evolutionary library (EL) in Datwarrior (DW) [message #1696] |
Tue, 09 August 2022 14:42  |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
I want to be able to copy a (any) structure from an existing table in DW into EL using a macro.
when I create a macro to do this it works fine.
However when I try to use this same macro on a new structure, it doesn't work. The macro just uses the previous structure and ignores the new one.
How do you get around this?
Ultimately I want to be able to run several EL's in a "chain" using a macro starting ideally from any structure to help towards creating a virtual library of new compounds.
Many thanks in advance
Jon
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1697 is a reply to message #1696] |
Tue, 09 August 2022 22:06   |
 nbehrnd
nbehrnd
Messages: 240
Registered: June 2019
|
Senior Member |
|
|
Dear Jon,
when generating an evolutionary library, DataWarrior offers (at least) twice to use a set of molecules from a .sdf or a .dwar as reference; in the screen photo below, they are marked by (A) and (B).
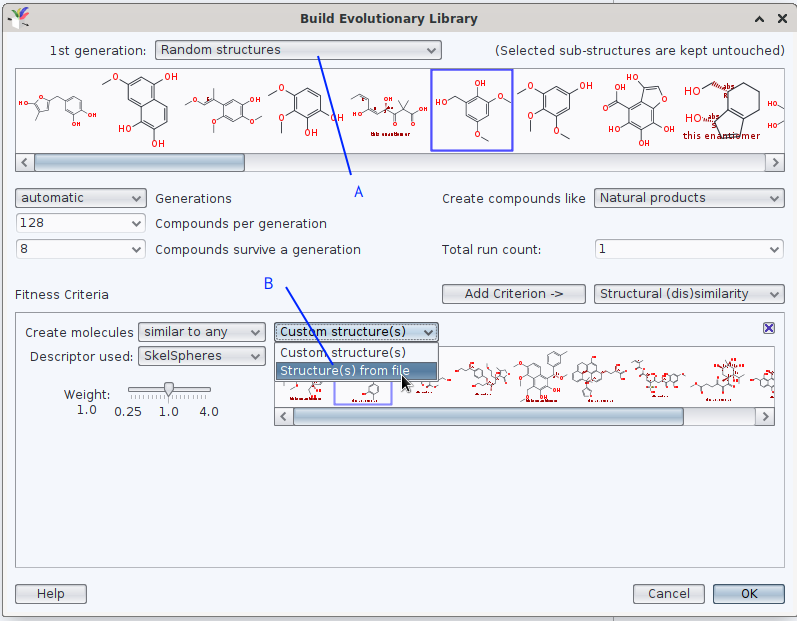
Should your question be understood in lines of «instead of molecules of an (of one) EL to be similar enough to a set of e.g., 20 molecules defined in box A, you would rather generate 20 EL where each of these reference molecules is a unique seed for an individual EL (more like an iteration)»?
Anyway, a .dwar and macro .dwam to represent a minimal working example (even if it currently stops after the first molecule of reference) to complement your question would be welcomed.
Norwid
[Updated on: Tue, 09 August 2022 22:19] Report message to a moderator |
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1700 is a reply to message #1697] |
Sun, 21 August 2022 19:31   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Hi Norwid and Thomas
Thanks for trying to answer the problem including the suggestion of using a file (which in this case did not solve the issue).
However, that's not quite what I had in mind so please see more information from the attached word document (with images).
Basically, it seems that you can't paste structures (or smiles) into an Evolutionary library window using a macro.
If I am correct - is there any way to change this as it would be really useful to have a chain of ELs running via a macro.
Many thanks in advance
|
|
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1702 is a reply to message #1701] |
Mon, 22 August 2022 17:57   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Here is the code from the macro that I used:
<macro name="Macro EL Help X">
<task name="selectView">
viewName=Table
</task>
<task name="buildEvolutionaryLibrary">
startSet=fb}@`@@YRYVum[ehrRkNBf@BjjjjJ`@@
survivalCount=8
kind=drugs
paramConfig0=structure similar SkelSphere s 100 fb}@`@@YRYVum[ehrRkNBf@BjjjjJ`@@
paramCount=1
generationCount=automatic
generationSize=128
</task>
<task name="selectWindow">
viewName=Evolutionary Library
</task>
<task name="selectView">
viewName=Table
</task>
<task name="sortRows">
column=Fitness
selectedFirst=false
descending=false
</task>
<task name="selectView">
viewName=Table
</task>
<task name="buildEvolutionaryLibrary">
startSet=fbm@`@@YrJIJEJJIRKTXxOdp@UUUUQQIQTMOP@
survivalCount=8
kind=drugs
paramConfig0=structure similar SkelSphere s 100 fbm@`@@YrJIJEJJIRKTXxOdp@UUUUQQIQTMOP@
paramCount=1
generationCount=automatic
generationSize=128
</task>
<task name="selectWindow">
viewName=Evolutionary Library
</task>
<task name="selectView">
viewName=Table
</task>
<task name="sortRows">
column=Fitness
selectedFirst=false
descending=false
</task>
<task name="selectView">
viewName=Table
</task>
<task name="sortRows">
column=Fitness
selectedFirst=false
descending=true
</task>
<task name="selectView">
viewName=Table
</task>
<task name="buildEvolutionaryLibrary">
startSet=fbmAB@L`dLbbRaRbbTbuFNCyL@EUUUTTRTUCSt@@
survivalCount=8
kind=drugs
paramConfig0=structure similar SkelSphere s 100 fbmAB@L`dLbbRaRbbTbuFNCyL@EUUUTTRTUCSt@@
paramCount=1
generationCount=automatic
generationSize=128
</task>
<task name="selectWindow">
viewName=Evolutionary Library
</task>
<task name="selectView">
viewName=Table
</task>
</macro>
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1703 is a reply to message #1702] |
Tue, 23 August 2022 21:58   |
 nbehrnd
nbehrnd
Messages: 240
Registered: June 2019
|
Senior Member |
|
|
Dear Jon,
while converting the macro you reported as text into a .dwam file (for convenience of further use attached below), I noticed multiple pairs where «a seed» reporting a molecules' structure in idcodes (the line starts with "startSet=") is followed by an other line beginning with "paramConfig0". Assuming the first line counted as line "1", in the present .dwam this pattern occurs thrice, i.e. for lines (6, 9), (29, 32), and (60, 63).
Simultaneously, I read on the second line of these a high SkelSphere threshold, 100, e.g.
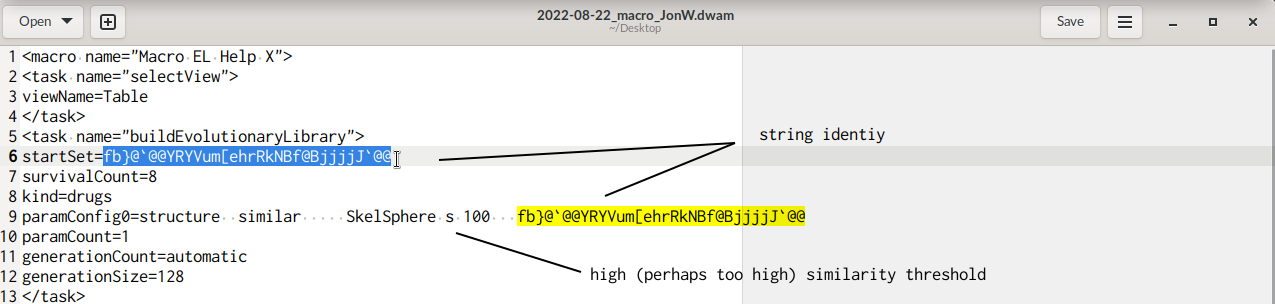
My speculation would be that this requested similarity might be too strict to yield multiple (new) molecules, perhaps especially because (based on your screen photos submitted) there only is one molecule to define the fitness of molecules to retain for a report / subsequent use.
Norwid
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1704 is a reply to message #1703] |
Tue, 23 August 2022 23:28   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Hi Norwid (and Thomas)
I have been using Notepad to edit the macro and found that DataWarrior saves the structure in the Macro as an Idcode.
The reason the Evolutionary Library appears three times as I am attempting to make it recursive and a minimum of two evolutionary libraries would be necessary, the third was just a demonstration of it not working. The SkelSpheres may be too high but in this example, the focus is on having the Macro "Copy and Paste" into the "startSet=...". The specifics of Weighting can be focused on at a later date.
Thanks for bringing to light the restriction of molecules I am generating and the fact that the target fitness is the same as the original. However, in the future, the objective is still to get "Copy and pasting" working inside the Macro's view of the evolutionary window.
Even if SkelSpheres structure similarity was at '60' and the parameters for the desired structure were different I would still have the same issue as I have tried configuring these in multiple ways before.
So any further help from you and Thomas on enabling copying and pasting in EL using a macro would be appreciated.
Thanks, Jon
|
|
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1707 is a reply to message #1706] |
Fri, 26 August 2022 15:40   |
 thomas
thomas
Messages: 744
Registered: June 2014
|
Senior Member |
|
|
Hi JonW,
One more remark: when after sorting a generated e-lib by fitness and before feeding those compounds into a new start generation, it would probably be useful to use just the best n compounds for the new start set. I didn't see a task to achieve that in you macro. One way of doing that would be to add row numbers after sorting and then applying a filter on the row number column e.g. from 1-32, then save visible rows as the file to be used for the next start set. Alternatively, you could select the table view, select all (visible) rows, copy view content (which in this case would be the entire TAB-delimited table) and use the selection as new 1st generation.
I noticed that a task to copy the n best molecule, providing n and the column to decide what is best, would be very handy here...
Hope this was more or less clear...
Thomas
[Updated on: Fri, 26 August 2022 15:42] Report message to a moderator |
|
|
|
|
|
|
|
|
|
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1722 is a reply to message #1717] |
Sun, 04 September 2022 19:57   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Dear Norwid
Many thanks for the link. A few questions:
1. Is the link you provided the same for any new updates/modifications that could be undertaken in the future or will a new link be created each time an update is made?
If it is the latter how do we know what the new link will be?
2. With regard to the download page at: https://openmolecules.org/datawarrior/download.html
This presumably has only the older "official version" ie DataWarrior V5.5.0 from April 2021 and does not have any updates since then- is that correct?
3. It may be useful to keep this older version in a separate folder in case one prefers some of the older settings (e.g. for running a specific macro for which a newer update might not function correctly). My question is - may an older version and an updated version on the same hard disc cause some stability issues or other problems?
Best regards
Jon
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1723 is a reply to message #1707] |
Mon, 05 September 2022 01:39   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Dear Thomas
Many thanks for the suggestions and for updating the software. I will try the changes this week and see how the macro performs.
Are there any published papers or further information on using the EL in DW apart from the online manual and a brief tutorial on youtube?
Best regards
Jon
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1724 is a reply to message #1723] |
Tue, 06 September 2022 08:05   |
 nbehrnd
nbehrnd
Messages: 240
Registered: June 2019
|
Senior Member |
|
|
Dear Jon,
the links following the penguin, the apple, the Windows symbol and before the «I have read and understood the disclaimer» which turn into green rectangles after your click: these are the links for the installers of DW for Linux, MacOS, and Windows respectively to provide the basic version of the program (at present, version 5.5.0) for major releases.
The address https://openmolecules.org/datawarrior/dw550win.zip only provides access to an archive to substitute the bits and bolts which changed since the release of the installer, version Windows. This sub set of data alone will not suffice to use DW. Without change of the address, the very same address is used for every update of the Windows version of DW. Speaking for the updates for MacOS/Linux I use, this either happens about once in 14 days (regular interval), or once a bug was identified and corrected. In the later case, uploads are updated as soon as possible (on occasion, multiple per day). This is why a keep a local copy of the installer, and of a few updater archives.
This allows to revert to an earlier minor version of DW should there be a need.[1] Then, deinstall DW completely, use the installer to install the major version 5.5.0 again, and eventually run the latest updater known to work well enough. Because the updater packages replace parts of DW's inner gear and I use only one installation directory for DW, there only is one version of DW installed (and most often working well). So far, there was no need for me to have multiple installations of DW (a minor release of June, and an other of August, etc.) to be present simultaneously. For one, the concurrent, simultaneous use of them would require additional guards to keep them separate in the computer's working memory (like in individual sandboxes,[2] or virtual machines[3]). For two, a user written macro should work regardless of the minor upgrade of DW used to write/record it, and irrespective if the underlying operating system was/is Windows, MacOS, Linux. Though only aware about two video recordings[4,5] I think Isabelle Giraud's workshop at RSC Open Chemical Sciences in 2020 set an other example of exchanging DW macros freely; they are meant to ease exchange among the interested. In case they do not work on more recent versions of DW, file a bug report; in case they don't work on an elder version of DW, perhaps the addressed functionality in DW wasn't yet implemented and you may get help here. Directory addresses -- if the macro uses them (many don't) -- may require an adjustment; yet because the macros are plain text, this requires a one-time edit already possible with a text editor.
Norwid
[1] example https://openmolecules.org/forum/index.php?t=msg&th=609&a mp;start=0&
[2] https://en.wikipedia.org/wiki/Sandbox_(software_development)
[3] https://en.wikipedia.org/wiki/Virtual_machine
[4] https://www.youtube.com/watch?v=mQCf9GakQW0
[5] https://www.youtube.com/watch?v=Is2hLqqSFvM
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1726 is a reply to message #1724] |
Thu, 08 September 2022 16:17   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Hi Norwid
If I don't use different versions of DW simultaneously -i.e., only use one at a time - can I still have two different versions on my PC?
Thanks for the links. These I already know. So my initial question remains: Are there any published papers or further information on using the EL (not using macros but the evolutionary library) in DW apart from the online manual and a brief tutorial on youtube?
I would like to understand the EL much better.
Many thanks
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1727 is a reply to message #1723] |
Thu, 08 September 2022 16:22   |
Jo W
Messages: 37
Registered: July 2021
|
Member |
|
|
Hi Thomas
Many thanks - the EL now works with the macro.
Thank you so much for your efforts.
With regard to my previous question and following Norwids response:
Are there any published papers or further information on using the EL in DW apart from the online manual and a brief tutorial on youtube?
Or failing that - is there a good paper on evolutionary libraries or even better some simple youtube videos that you know of that would help, ideally, specifically in understanding how EL works in DW?
Best regards
|
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1728 is a reply to message #1727] |
Thu, 08 September 2022 20:15   |
 nbehrnd
nbehrnd
Messages: 240
Registered: June 2019
|
Senior Member |
|
|
Dear Jon,
the safest approach is to run the installer and the update A for version A. And if you want to check version B, to 1) deinstall DW, 2) run the installer, and 3) the updater for version B). This is by precaution because I do not know if updater A and B contain the same libraries by name the updated libraries of updater A may simply exchanged by those of updater B (like different tires on a car), .or. if they introduce changes in the other components of DW.
As for external literature references of evolutionary libraries .and. DataWarrior, my answer is no, I do not have a good i.e., tutorial-like literature reference at hand. Ningsih et al.[1] describe they used it, however address those who already are in the know:
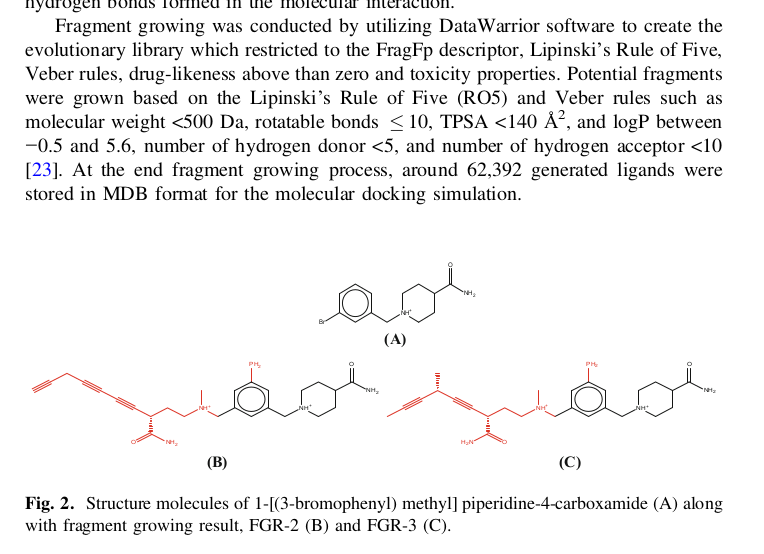
(loc. cit. p. 18)
[1] Ningsih, E.G., Hidayat, M.F., Tambunan, U.S.F. (2019). Fragment-Based Drug Design to Discover Novel Inhibitor of Dipeptidyl Peptidase-4 (DPP-4) as a Potential Drug for Type 2 Diabetes Therapy. In: Rojas, I., Valenzuela, O., Rojas, F., Ortuño, F. (eds) Bioinformatics and Biomedical Engineering. IWBBIO 2019. Lecture Notes in Computer Science(), vol 11465. Springer, Cham. https://doi.org/10.1007/978-3-030-17938-0_2 pages 14–24 (paywall).
[Updated on: Thu, 08 September 2022 22:09] Report message to a moderator |
|
|
|
| Re: Macro for evolutionary library (EL) in Datwarrior (DW) [message #1731 is a reply to message #1728] |
Sun, 11 September 2022 15:33   |
 thomas
thomas
Messages: 744
Registered: June 2014
|
Senior Member |
|
|
Dear Norwid,
many thanks for answering all the questions already...
Dear Jon,
let me quickly add some comments to two questions:
If you want to keep running the newest version: Once install the original 5.5.0 version from 2021 (which you have already done). Once in a which download the dw550win.zip (Mac or Linux dw550x.zip) using the small print link from the official datawarrior download page. Unpack the file to obtain a few replacement files. Make a save copy of the respective original files with the same names and move the files from the zip archive to the DataWarrior installation folder, where they replace the original files. To move to an earlier version you can just copy an earlier set of files or the saved original files back to the DataWarrior installation folder.
I am aware that the explanation of how to use the evolutionary library and how it works is not well represented in the manual. Especially, the new fitness criteria PheSA and docking have a lot of potential. The best thing to understand what it really does one might look a the source code, but admittedly, that is not for everybody and it still takes some time even for a person being fluent in Java. I also realized that the current algorithm could be improved by reusing fragments of previous hi-ranking structures. The current principle is rather simple: apply simple modifications to the best structures of the previous generation, rank them and use the best ranking molecules for further modifications. Modifications are selected randomly from a complete list of available modifications. The random selection is completely random: modifications are the more likely, the more the product is drug-like (or natural product like). This ensures that created molecules make sense in this regard. The central Java class that performs the changes is called Mutator and can be found here. It lists the kind of changes possible and also contains the few lines of code to actually perform the change in the molecule. The rest is basically how to calculate fitness, which is a weighted sum of user selected parameters. The more complex are Flexophore similarity (DOI:10.1021/ci700359j), PheSA (paper will come) and Docking.
Hope this is somewhat useful...
Thomas
|
|
|
|
|
|
|
|
Goto Forum:
Current Time: Fri Apr 03 10:36:59 CEST 2026
Total time taken to generate the page: 0.00774 seconds
|
