| Re: Visualising SMILES string [message #1261 is a reply to message #1258] |
Tue, 23 March 2021 16:38   |
 thomas
thomas
Messages: 747
Registered: June 2014
|
Senior Member |
|
|
Dear Angus,
I have extended the SmilesParser within OpenChemLib and used by DataWarrior to support '*' and '?' as pseudo atom symbols. Both symbols can be used inside or outside square brackets. While '*' is inline with the opensmiles.org standard, '?' is not. Nevertheless, because of the missing distinction in the smiles syntax between query fragments and full molecules, I believe that '?' makes sense.
'*' creates a wild card atom, which is an atom query feature and only allowed in query fragments. Therefore, a smiles containing a '*' is atomatically perceived as a query structure rather than a full molecule. Its free valences are not considered as being filled with hydrogen.
'?' is converted into an atomicNo=0, which DataWarrior uses as attachment point, e.g. in R-groups after a SAR deconvolution. These atoms are meant to not exist. Their sole purpose is to carry the bond sticking out of the R-group.
If you paste the these 6 rows of smiles:
C1CCC([*:1])CC1O
C1CCC([*])CC1O
C1CCC(*)CC1O
C1CCC([?:1])CC1O
C1CCC([?])CC1O
C1CCC(?)CC1O
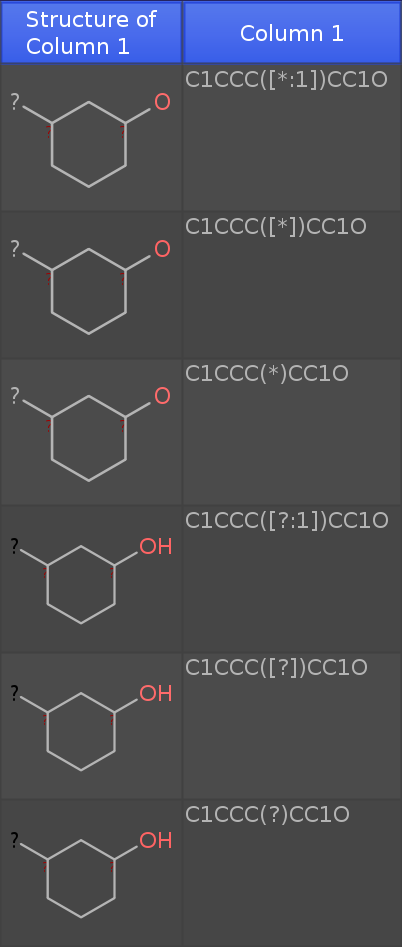
into a new DataWarrior Window (newest dev release), then you get the following table:
Note that I have added an oxygen atom. If you see an 'H' at the oxygen, the structure is considered a molecule. If not, the structure is a query fragment with open valences.
-
 Attachment: t.png
Attachment: t.png
(Size: 41.90KB, Downloaded 1773 times)
|
|
|
|
