| suggest: native .pdf export [message #897] |
Mon, 11 May 2020 12:29  |
 nbehrnd
nbehrnd
Messages: 233
Registered: June 2019
|
Senior Member |
|
|
Based on a .smi derived from elsewhere,[1] which was converted with openbabel
into a .sdf I let DataWarrior screen the structures for a few characteristics.
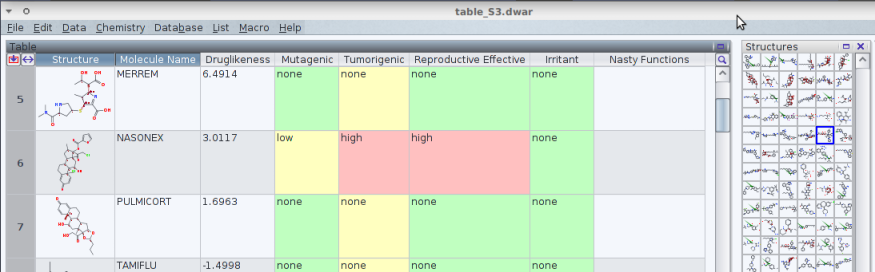
Aiming to share it with one who does not use DataWarrior, I miss a built-in
function to print the table as a .pdf file as a vehicle of discussion.
This could be useful, especially if -- as here -- the cell's background colour
has a significance.
So far, I printed the table via an installed HP printer as a postscript into
a file (of very large file size) and converted it with ps2pdf into a .pdf
like the example file attached. At page breaks, however, lines seem to be
broken (table headings). If exported directly from DW as .pdf, their file size
could be considerably smaller than now (containing an image in the .pdf as a
container) and possibly retain a searchable text-layer.
[1] https://pubs.acs.org/doi/suppl/10.1021/jm301008n/suppl_file/ jm301008n_si_002.xlsx
-
 Attachment: table_S3.png
Attachment: table_S3.png
(Size: 95.75KB, Downloaded 1195 times)
-
 Attachment: table_S3.dwar
Attachment: table_S3.dwar
(Size: 22.49KB, Downloaded 538 times)
-
 Attachment: table_S3.pdf
Attachment: table_S3.pdf
(Size: 1.81MB, Downloaded 565 times)
|
|
|
|
|
|
| Re: suggest: native .pdf export [message #903 is a reply to message #898] |
Tue, 12 May 2020 16:03   |
 nbehrnd
nbehrnd
Messages: 233
Registered: June 2019
|
Senior Member |
|
|
It was possible to replicate the indicated method using the cups pdf printer.
As a closing comment:
Still interested to benefit more from the vector format I wrote a Python script
that reads some of DW's .dwar file content and the retained list of SMILES strings,
calls openabel to visualize the structures, and puts all in an .xlsx file. The
manual work then left was to open this file in LibreOffice Calc, to adjust the
images' sizes to fit the cell size, to apply conditional cell background colors
and to save it as .ods. The file size of the then exported .pdf is slightly less
than half of the one printing from DW with cups while still offering a searchable,
crisply printed text layer, too.
-
 Attachment: table_S3.smi
Attachment: table_S3.smi
(Size: 7.79KB, Downloaded 507 times)
-
 Attachment: table_S3.dwar
Attachment: table_S3.dwar
(Size: 22.49KB, Downloaded 518 times)
-
 Attachment: spreadsheet_test.py
Attachment: spreadsheet_test.py
(Size: 4.35KB, Downloaded 585 times)
-
 Attachment: test.ods
Attachment: test.ods
(Size: 1.08MB, Downloaded 524 times)
-
 Attachment: test.pdf
Attachment: test.pdf
(Size: 850.78KB, Downloaded 591 times)
|
|
|
|
|
|
| Re: suggest: native .pdf export [message #914 is a reply to message #907] |
Fri, 22 May 2020 00:25  |
 thomas
thomas
Messages: 731
Registered: June 2014
|
Senior Member |
|
|
DataWarrior uses TAB or comma delimited text files. The smi File only contains spaces between Smiles and Name. If you replace all SPACEs by a TABs in any Text-Editor before pasting into DataWarrior, you will correctly get three columns: Structure, Smiles, and Name
[Updated on: Fri, 22 May 2020 00:26] Report message to a moderator |
|
|
|
